Leakly
Leakly: Leakage checks for any machine-learning pipeline
Leakly uses label permutation to test whether a machine-learning pipeline performs above chance when no true signal is present.
Above-chance performance after permutation may indicate leakage from preprocessing, feature selection, tuning, or another step of the pipeline.
How it works
- Permute labels to remove the real feature-label association.
- Run the full pipeline exactly as in the original analysis.
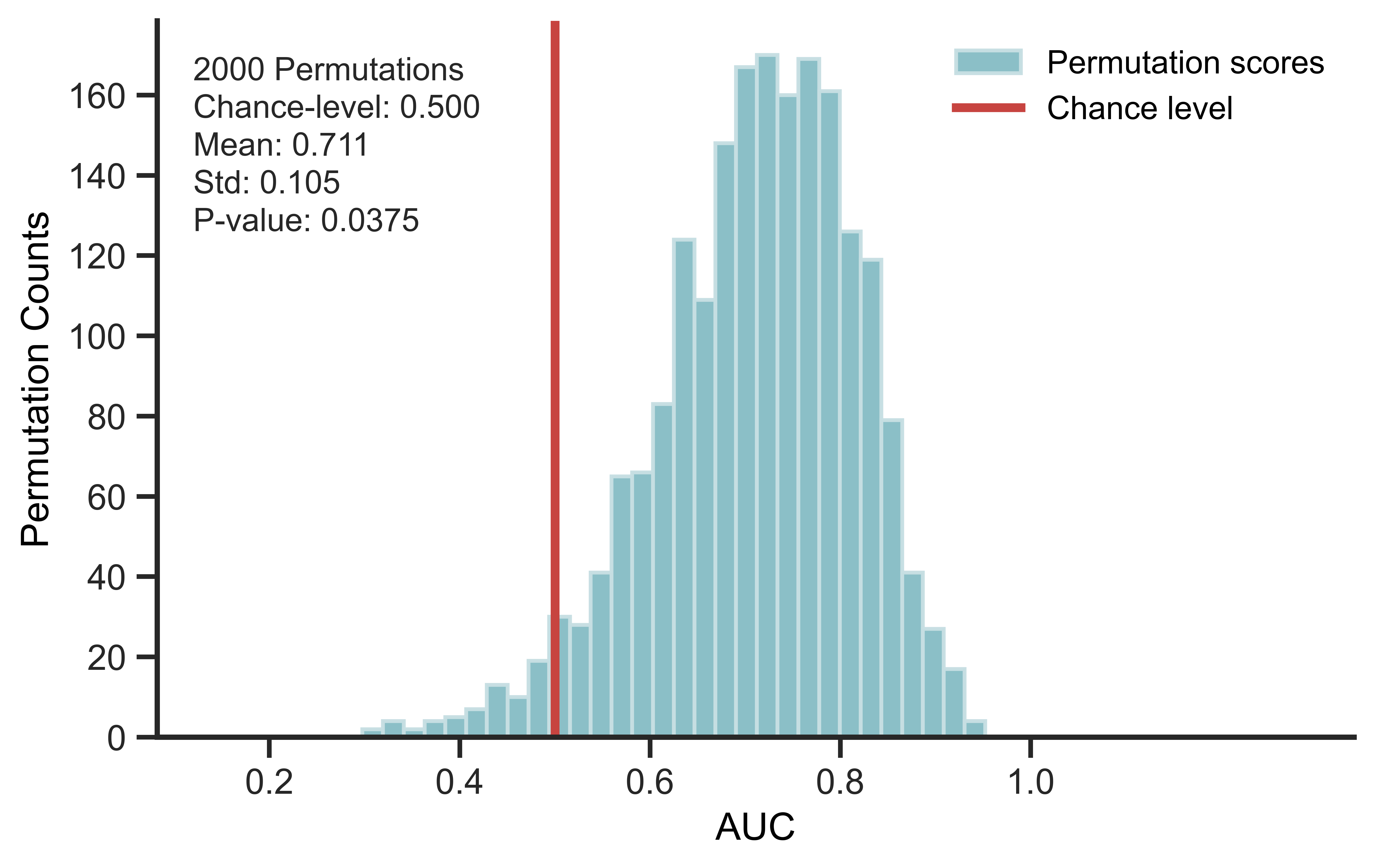
- Compare the permuted score distribution with chance level.
- Above-chance permuted performance suggests possible leakage.
Leakly includes example configurations for a leaky pipeline and a non-leaky pipeline so users can inspect the effect directly.
Install
pip install Leakly
Quick Start on Colab: 
Key Python snippet
from leakly import (
MLPipeline,
SummaryPlotter,
load_example_leakage_config,
permute_label)
scores = []
for seed in range(100):
permuted_y = permute_label(y, random_state=seed)
score = (
# Replace with any user-defined pipeline
MLPipeline(
X,
permuted_y,
covariates=covariates,
config=load_example_leakage_config(),
).fit()
).evaluate()
scores.append(score)
SummaryPlotter(scores, chance_level=0.5).plot()
FAQ
Can Leakly check my own pipeline?
Yes. Leakly can evaluate any pipeline that takes X, y,
optional covariates, and returns a test score.
The key is to run the full pipeline exactly as in the real analysis,
including preprocessing, feature selection, tuning, and evaluation.
Why can a leaky pipeline score well on permuted labels?
If leakage occurs, information from test samples can enter the analysis before the train/test split or outside the cross-validation loop. Common sources include feature selection, scaling, imputation, covariate adjustment, dimensionality reduction, or hyperparameter tuning performed on all samples.
In high-dimensional data such as omics and neuroimaging, random features can appear predictive by chance. If a pipeline can retain these spurious patterns, it may perform above chance even after labels are permuted.
How many permutations should I run?
Use 100 for a quick check. Use 1,000 or more for publication-level evidence.
License
MIT. See LICENSE.

